FlyWire is a community data resource containing whole brain connectome of the female adult fruit fly ( FAFB dataset). The central brain has been fully proofread by human experts and as of Oct 2022, the whole brain is 85% proofread. The community has added over 33,000 labels to the volume.
The FlyWire interface:
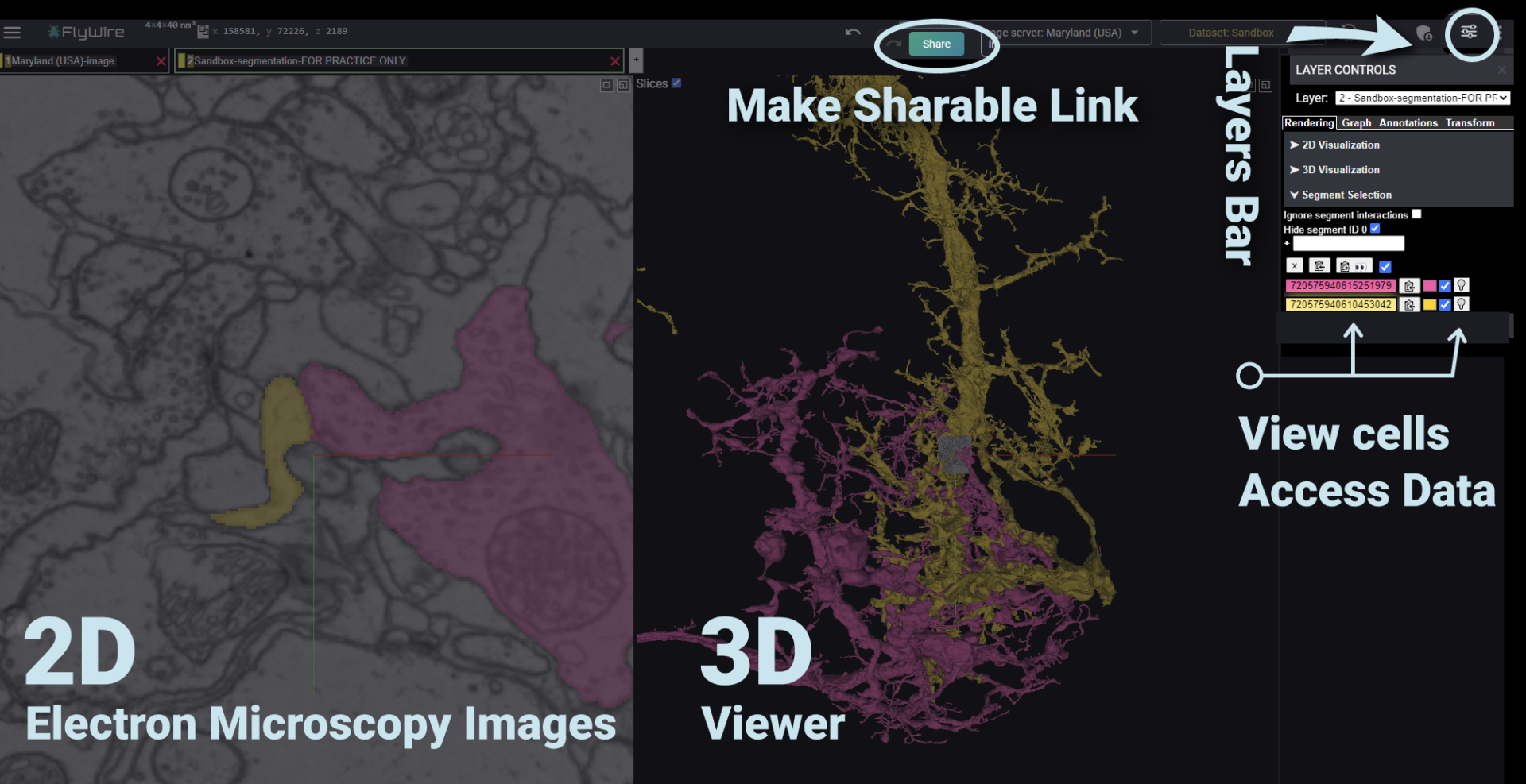
After you sign into FlyWire, navigate with the following controls:
2D (EM) or 3D Navigation
- Add or remove segments – Double click
- Zoom – CTRL + scroll
- Jump to location – Right click area of interest
- Undo action – CTRL + Z
Navigating 3D space
- Rotate – Click + drag
Navigating 2D (EM) space
- Pan – Click + drag
- Step through slices – scroll
For a deeper dive into FlyWire controls go here.
Finding neurons
If you know of a neuron in the FlyCircuit or the Hemibrain datasets, copy its ID number from that dataset and paste it into the FlyWire Gateway to see a point cloud of its neurites in FlyWire. Click neurons near those points in FlyWire to look for a match.
If you’re looking for a neuron you know of from the Virtual Fly Brain (VFB) site, select one of its images there. Download an .SWC file, if one is available (usually for neurons that came from EM), and upload it into the FlyWire Gateway, selecting the “JRC2018U” template in the Gateway. As above, the resulting point cloud in FlyWire can be used to guide a search of nearby neurons for a match. If an .SWC is not available, download a .NRRD file, and create an .SWC yourself.
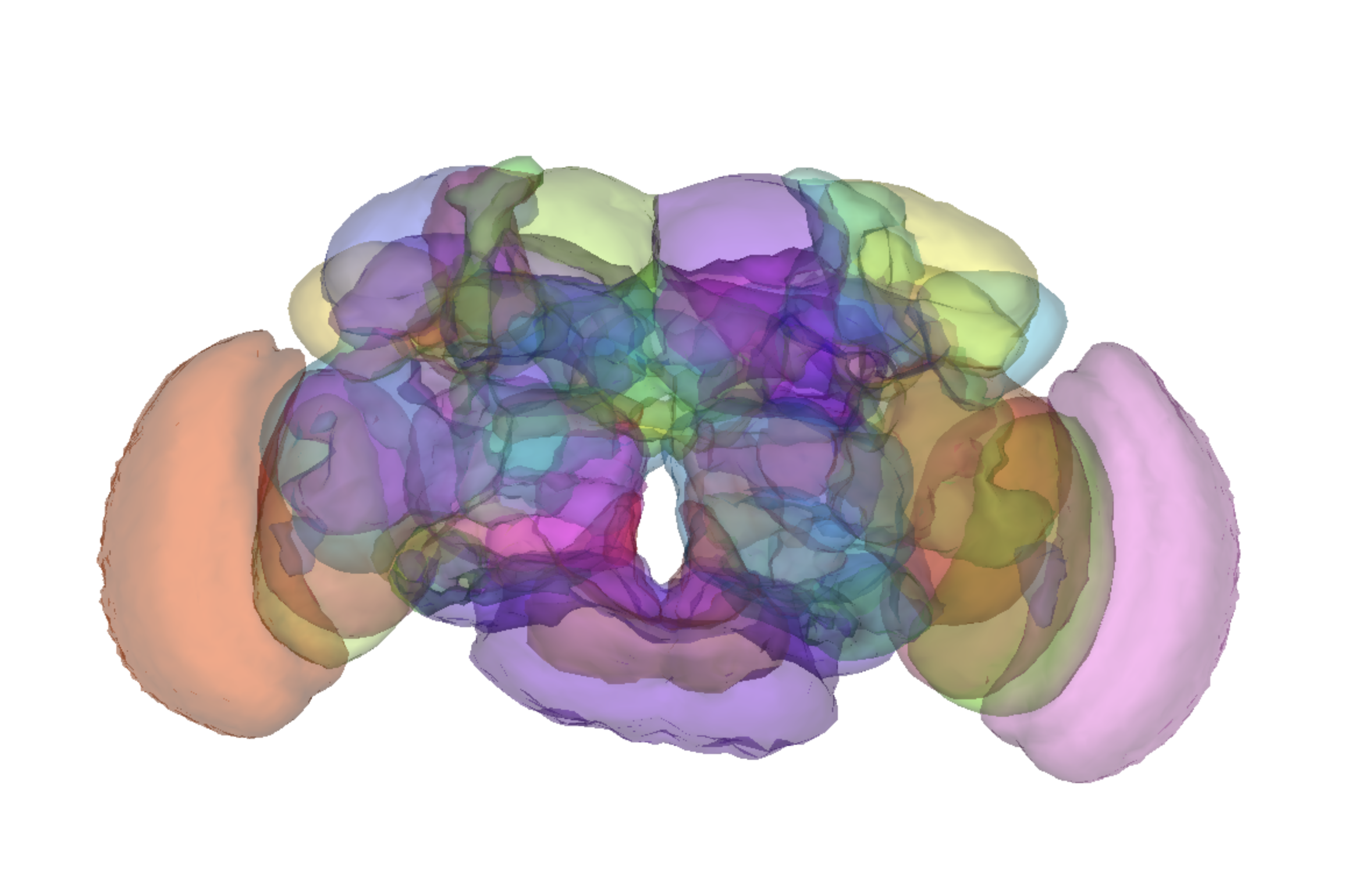
The Jefferis lab and Google overlaid a map of brain neuropil regions into the FlyWire. To see the neuropil masks alongside neurons you’ve been proofreading, use this link.
Annotating Neurons
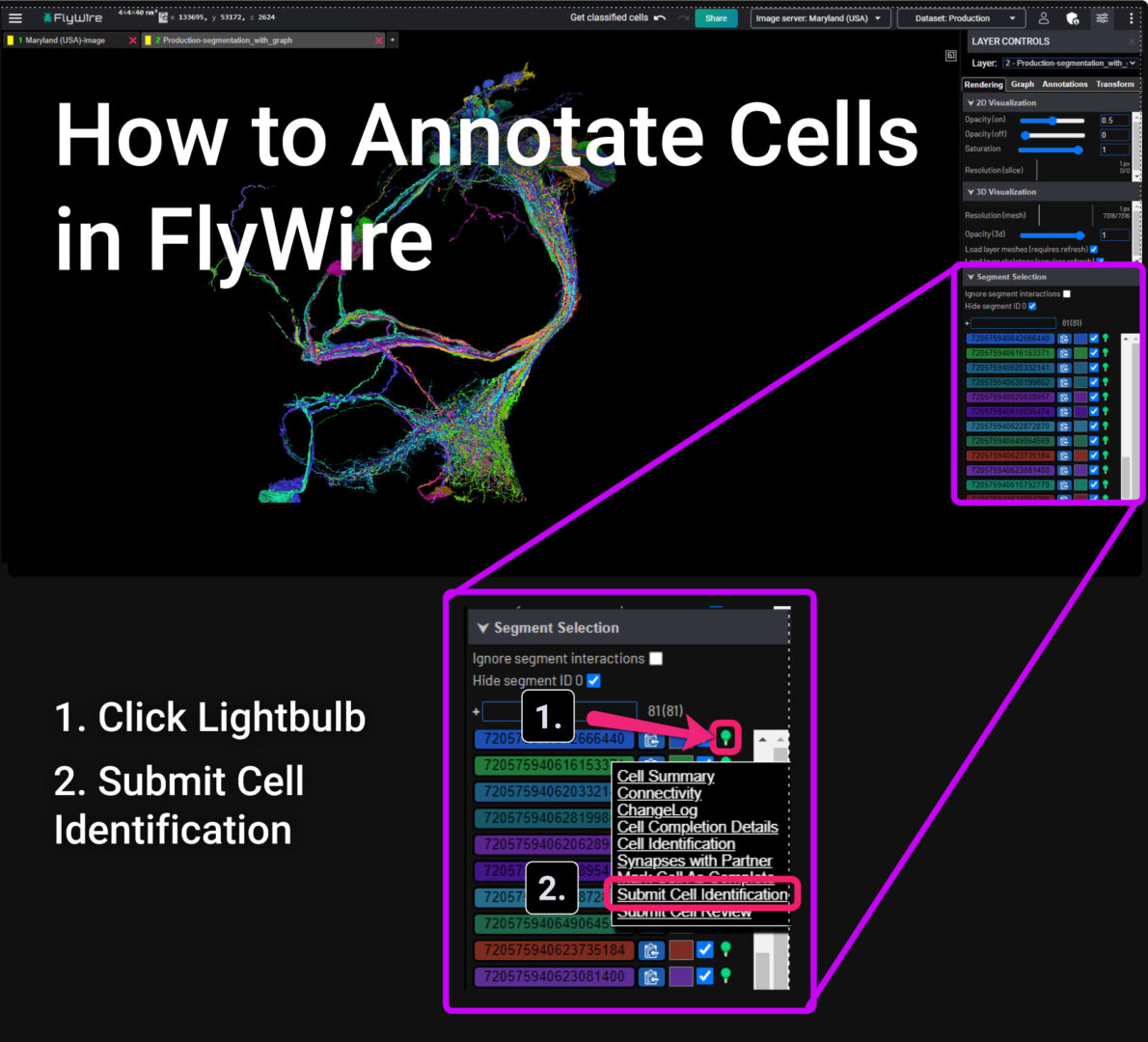

Annotation Layers
- Add an annotation layer by right-clicking the plus button ➕ to the right of your active layers.
- Right-click the annotation layer to select it
- Click an annotation type (annotate point suggested). Selected annotation type will turn green.
- CTRL + Click in the 2D (EM) or 3D to place your annotation
One response to “How to Use FlyWire”
How does the FlyWire community data resource, featuring the whole brain connectome of the female adult fruit fly, benefit research? Regard Telkom University